Monday May 26, 2025
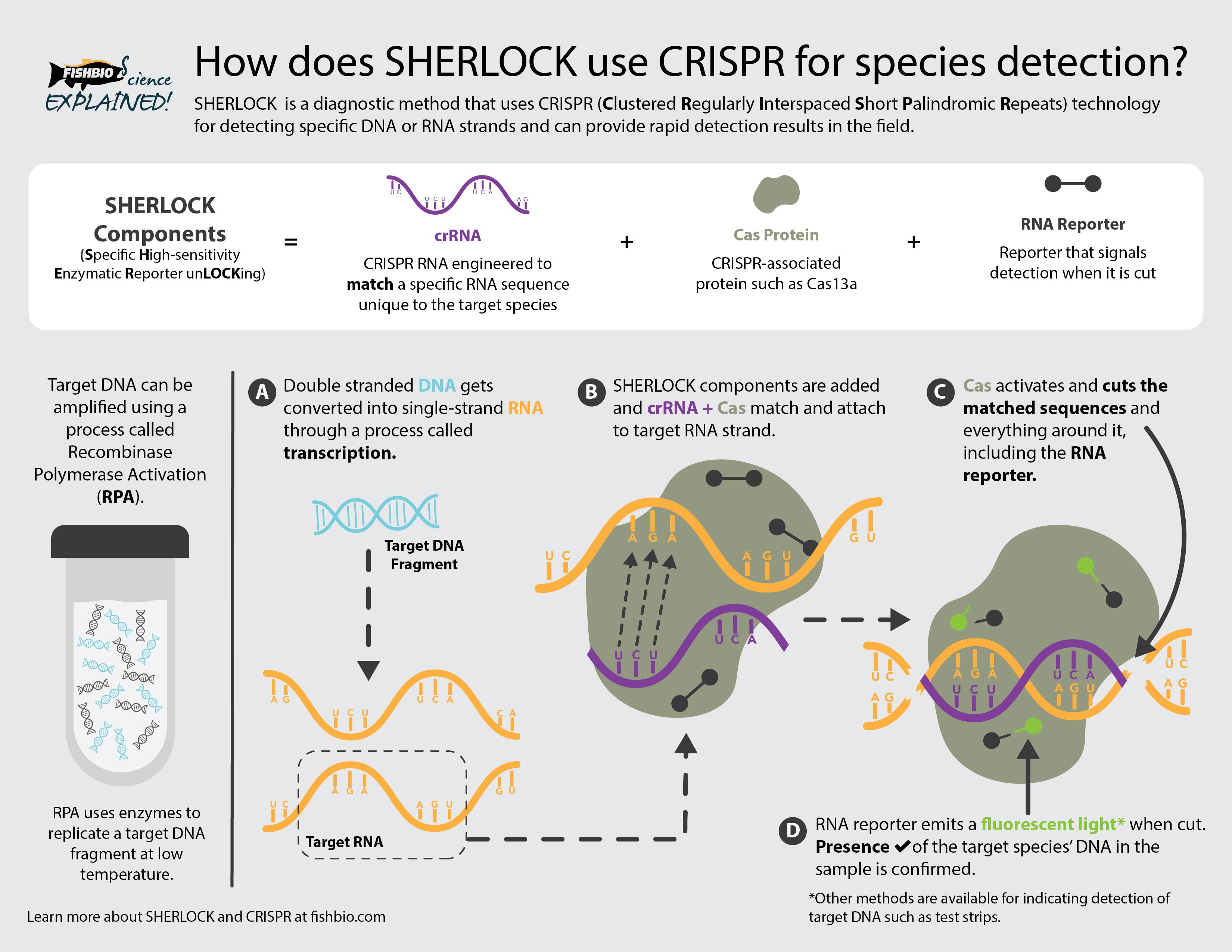
While it sounds like an air fryer setting, CRISPR-based (pronounced “crisper”) genetic methods have revolutionized biomedical, agricultural, and wildlife and fisheries sciences. The CRISPR/Cas system’s (Cas is short for CRISPR-associated protein) ability to locate, cut, and modify specific genes has made it useful for various applications, including treating genetic diseases and improving the nutritional content or pest resistance of some agricultural crops. CRISPR methods can also be used for diagnostic purposes to identify viral or bacterial infections and even species identification. Specific High-sensitivity Enzymatic Reporter UnLOCKing – or, SHERLOCK – is a CRISPR-based method utilizing slightly different Cas proteins to detect species-specific DNA. Researchers use SHERLOCK to distinguish between similar-looking smelt species and Chinook salmon (Oncorhynchus tshawytscha) run types and are expanding this method into environmental DNA (eDNA) studies. But how does SHERLOCK work?
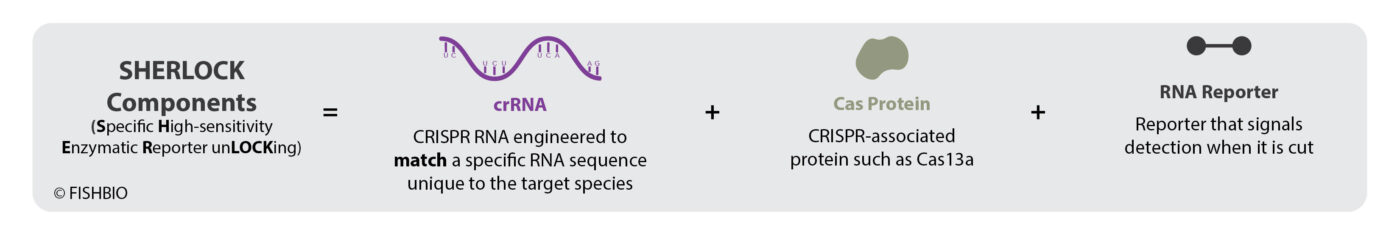
On a molecular level, CRISPR and SHERLOCK technology can seem complex, but it can be broken down into a few key parts. The CRISPR system has two main components: 1) Cas proteins, the defense mechanisms for some bacteria, whose job it is to chop up, or cleave, DNA or RNA at specific locations, and 2) CRISPR RNAs (crRNAs) that bind with Cas proteins, allowing them to recognize those specific locations.
There is also a reporter molecule in the SHERLOCK method that provides a signal if it gets cleaved. This RNA reporter will only get chopped up if the SHERLOCK Cas protein (Cas13a) finds the specific RNA sequence of interest.
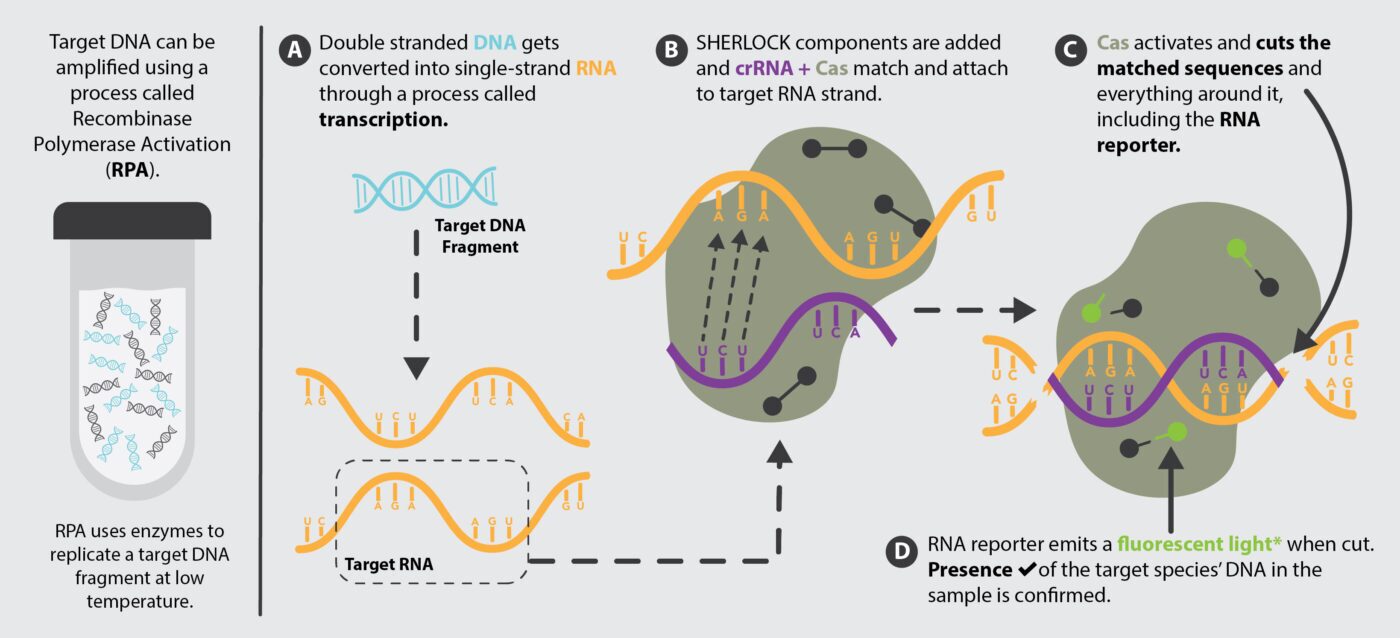
To make this all work, scientists first find segments of DNA whose sequence is unique to the species of interest (target DNA). For example, mitochondrial DNA is commonly used for fish species identification. Next, scientists have Cas13a proteins and crRNAs engineered to recognize the exact target RNA sequence corresponding to the target DNA (i.e., the RNA sequence UCUU corresponds to the RNA sequence AGAA). Then, they combine sample DNA with the SHERLOCK components in one or more chemical reactions providing ideal conditions to: 1) replicate target DNA if it is within the sample (making more copies makes it easier to detect), 2) convert the DNA to RNA (a process called transcription), and 3) allow SHERLOCK components to recognize and bind to target RNA. If the target RNA is present, then the reporter molecule gets cleaved and will signal the presence of the target DNA.
Sample DNA can be acquired by different methods, including tissue samples from an individual needing identification, mucus swabs, and eDNA (where fish handling is not required). Tissue samples are subjected to chemical digestion to release DNA from the tissue cells. Since DNA in mucus is not contained within excess cellular material, as in tissue, it requires less digestion and can more readily be used for the SHERLOCK method.
There are also different methods to amplify target DNA for SHERLOCK. The traditional approach, known as polymerase chain reaction (PCR), uses heat to separate DNA strands and cooling to allow enzymes to make copies of target DNA. This process is performed using a laboratory technique called quantitative PCR (qPCR). More recent advancements in DNA amplification are methods that do not require cycling between hot and cool temperatures and can be performed at a single temperature (known as isothermal amplification). One method that works with SHERLOCK is called recombinase polymerase amplification (RPA), which leverages enzymes to separate double-stranded DNA and replicate target DNA at temperatures just warmer than room temperature.
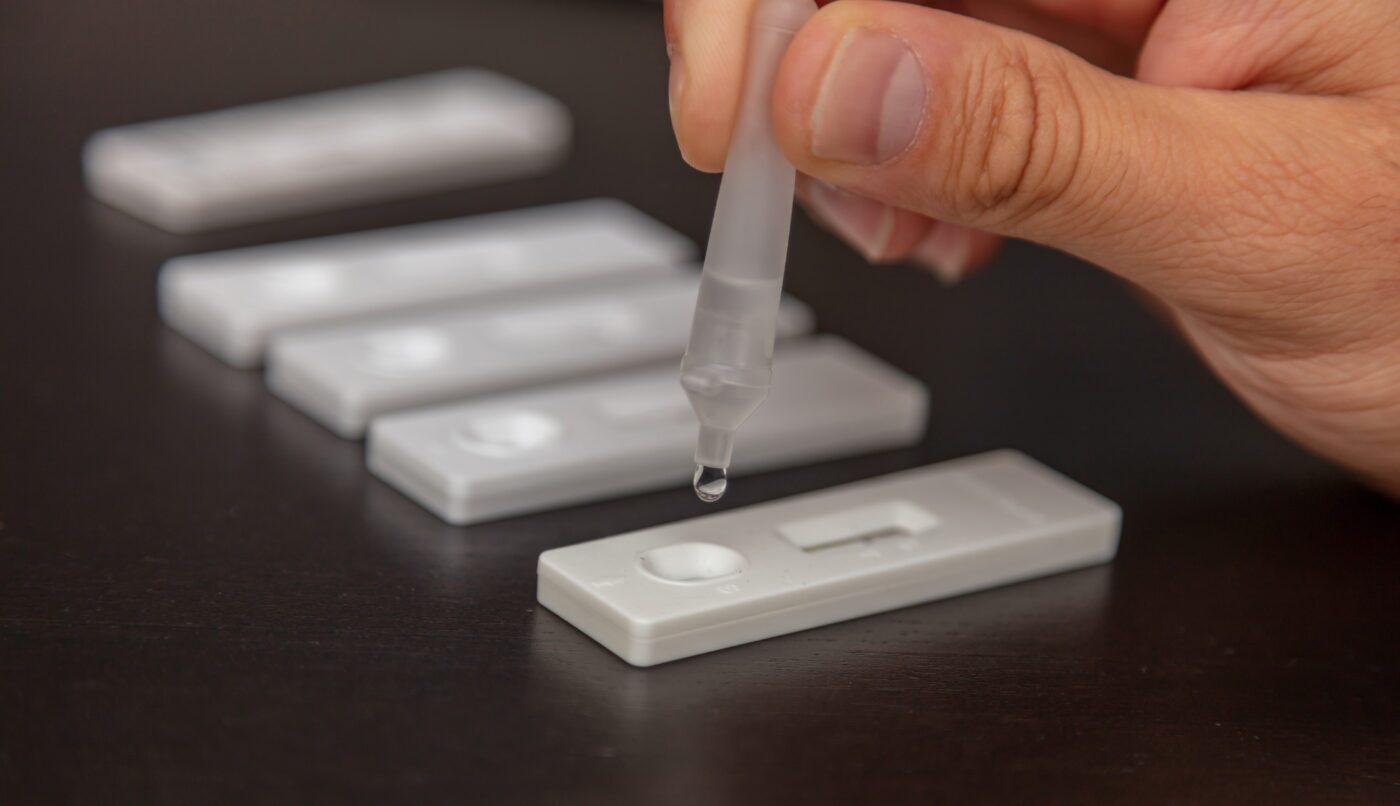

Just as there are options for target DNA amplification, there are options for target DNA detection (the last step of SHERLOCK requiring the RNA reporter). Some RNA reporters have a fluorescent tag that emits a tiny burst of light when the reporter is cut. Light from these reporters can be detected and measured with an instrument. There are also other RNA reporters that can bind to different areas on a test strip if they are chopped. Assessing detection for these is similar to reading a positive or negative result from an at-home Covid test.
The CRISPR-based method SHERLOCK is a game changer in genetic species identification. Continued development will take the laboratory to the field, assisting in differentiating between visually similar species or providing rapid field-based detection of invasive, rare, or endangered species in eDNA assessments.
This post was featured in our weekly e-newsletter, the Fish Report. You can subscribe to the Fish Report here.

